ChromHMM
Chromatin state discovery and characterization
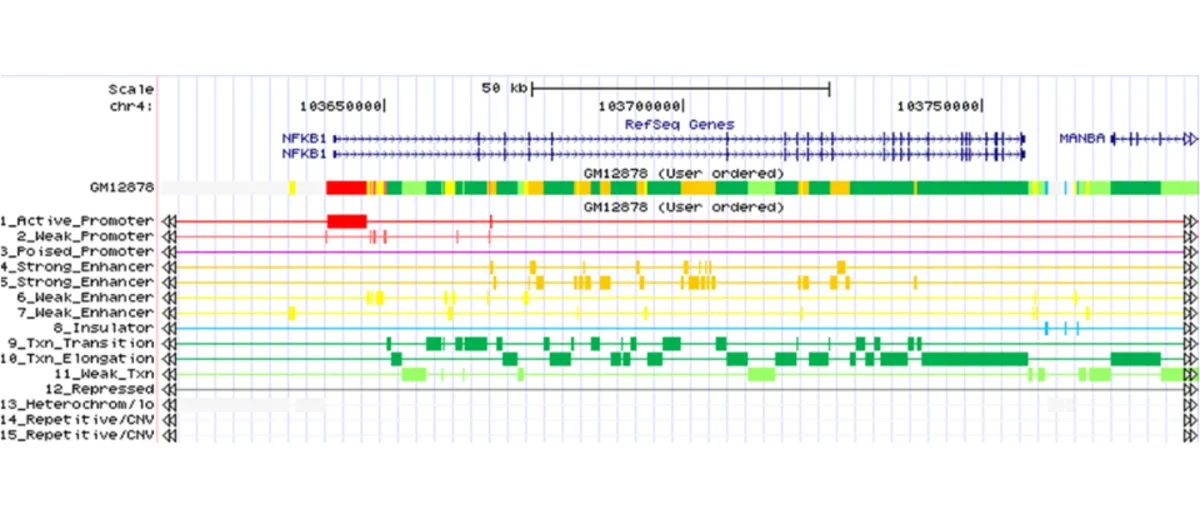
ChromHMM is software for learning and characterizing chromatin states. ChromHMM can integrate multiple chromatin datasets such as ChIP-seq data of various histone modifications to discover de novo the major re-occuring combinatorial and spatial patterns of marks. ChromHMM is based on a multivariate Hidden Markov Model that explicitly models the presence or absence of each chromatin mark. The resulting model can then be used to systematically annotate a genome in one or more cell types. By automatically computing state enrichments for large-scale functional and annotation datasets ChromHMM facilitates the biological characterization of each state. ChromHMM also produces files with genome-wide maps of chromatin state annotations that can be directly visualized in a genome browser.
Access the main ChromHMM website
Citation:
Ernst J, Kellis M.
ChromHMM: automating chromatin-state discovery and characterization.
Nature Methods, 9:215-216, 2012.