CNEP
Constrained Non-Exonic Predictor
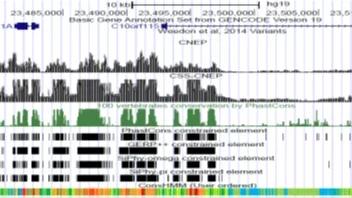
CNEP is software for predicting constrained-non exonic bases from large scale epigenomic and transcription factor binding data.
Access the CNEP project on GitHub
Citation:
Grujic O, Phung TN, Kwon SB, Arneson A, Lee Y, Lohmueller KE, Ernst J.
Identification and characterization of constrained non-exonic bases lacking predictive epigenomic and transcription factor binding annotations.
Nature Communications, 11:6168, 2020.